Rutgers Scientists Discover Molecules that Show Promise for New Anti-Flu Medicines
Chemicals block ability of flu virus to replicate in cells; goal is to develop medicines that fight much-feared pandemic influenza outbreaks
NEW BRUNSWICK, N.J. – A new way to attack flu viruses is taking shape in laboratories at Rutgers University, where scientists have identified chemical agents that block the virus’s ability to replicate itself in cell culture.

These novel compounds show promise for a new class of antiviral medicines to fight much-feared pandemic influenzas such as the looming “bird flu” threats caused by the H5N1 influenza A virus and the new H7N9 virus responsible for a 2013 outbreak in China.
Timely production of a vaccine is difficult when a pandemic flu strikes. A viable alternative is to treat with drugs.
“Right now there’s really only one effective oral drug for treating influenza,” said Eddy Arnold, Board of Governors Professor of chemistry and chemical biology in the School of Arts and Sciences at Rutgers and a member of the Center for Advanced Biotechnology and Medicine. And just as bacteria develop resistance to antibiotics, Arnold notes that some flu strains have developed resistance to Tamiflu, the sole orally available anti-flu drug.
Arnold and his collaborators have been working to create drugs beyond Tamiflu, especially ones that target different parts of the virus, using an approach that helped in the development of powerful anti-AIDS drugs. By synthesizing chemical compounds that bind to metal ions in a viral enzyme, the researchers found they could halt that enzyme’s ability to activate a key step in the virus’s replication process.
In Arnold’s words, his team’s compounds “really gum up” the targeted enzyme of influenza virus.
“We’re at a key proof of principle stage right now,” he said. “It’s not trivial to go from this point to actually delivering a drug, but we’re optimistic – this class of inhibitors has all the right characteristics.”
Rutgers’ search for these binding compounds relies on technology that reveals the structure of this enzyme in extremely fine detail. Researchers Joseph Bauman and Kalyan Das first produced high-resolution images of an H1N1 flu enzyme, and Bauman and postdoctoral researcher Disha Patel screened 800 small molecule fragments for binding.
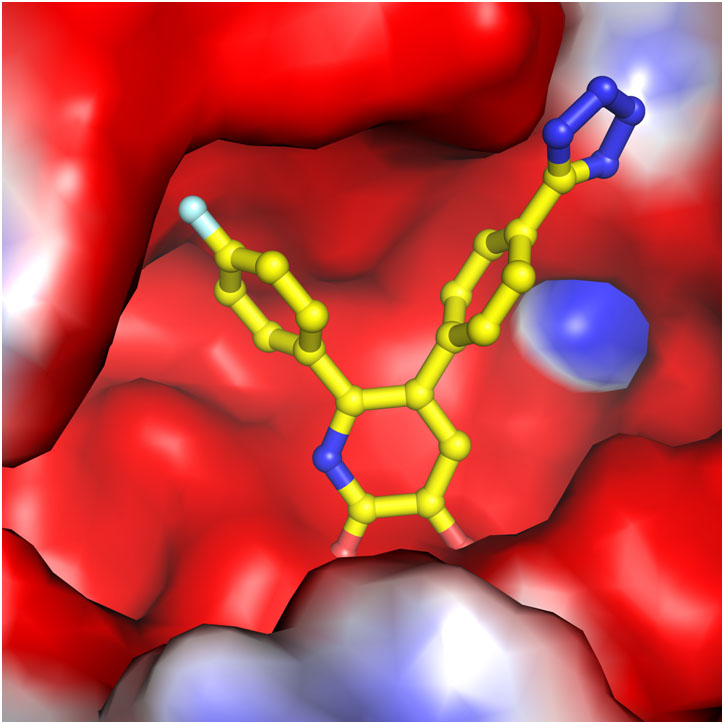
The researchers in Arnold’s lab worked with Edmond LaVoie, professor and chair of medicinal chemistry in the Ernest Mario School of Pharmacy, to modify those compounds, making them more potent and selective in blocking the flu enzyme’s activity. Working with virologist Luis Martinez-Sobrido at the University of Rochester, they were able to detect antiviral activity of the compounds in cells.
The enzyme that the scientists are attacking is especially crafty, Arnold noted, because it steals material from human cells to disguise the invading flu virus in a process called “cap-snatching.” These “caps” are a small chemical structure that prime the process for reading genetic information.“What we’re doing by blocking or inhibiting this enzyme is to interefere with flu’s ability to disguise itself,” he said.
Arnold cited research by universities and pharmaceutical companies nearly two decades ago that took this approach, but initially the technology to obtain high-resolution images of the influenza protein wasn’t available. One pharmaceutical company, Merck, later applied the approach of targeting metal-ion containing active sites in the HIV enzyme integrase and developed a highly successful anti-AIDS drug.
“It’s truly remarkable what they did, and we’re trying to pursue similar logic with influenza,” said Arnold.
The researchers have recently published their findings in the American Chemical Society journal ACS Chemical Biology (http://pubs.acs.org/doi/abs/10.1021/cb400400j). Some of the work was funded by the National Institutes of Health. Two additional publications in the journals Bioorganic Medicinal Chemistry and ACS Medicinal Chemistry Letters have described LaVoie’s synthetic medicinal chemistry used to make the new anti-flu agents and the observed structure-activity relationships.